Now I turned this blog post into an arXiv preprint: https://arxiv.org/abs/2007.14386
Here, I will discuss some technical issues of compartmental models in general, and the SIR model in particular, on temporal networks. These are things that feel a bit too off-topic to even bother readers of papers, but everyone into network epidemiology needs to consider them at some point. Maybe I should write a tutorial paper about it (the intro chapter in Naoki Masuda’s and my book covers some of these issues), in the meanwhile, here we go:
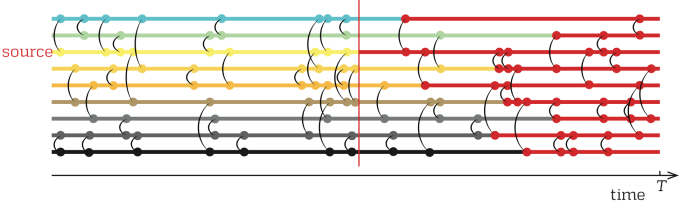
Assume that we know a sequence of contacts describing the interactions within a population. Assume this sequence takes the form of a list of triples (i,j,t) that tells us that nodes i and j are in contact at a (discrete) time t. Then we assume a disease outbreak could be described by the SIR model over these contacts. I.e., we can divide the population into susceptible (S), infectious (I), and recovered (R) individuals. If one node of a contact is S and the other I, then S becomes I with probability β. This is the basic setup. Then one needs to choose between several details of the model. Over the years, I have been into epidemic modeling, I have sometimes flipped between different choices. Most choices are not right or wrong, but some are more right than others. Here is what I do these days, and why.
- Start the outbreak at one node. This is for the very fundamental reason that epidemiologists are usually concerned with the outbreak of one strain arising from a mutation in one host, or entering the population from one external (zoonotic) interaction. If one starts the epidemic at different sources, one would be assuming that there had been some spreading outside of the network. Thus that one is modeling an open system. Therefore one needs to model the possible influx of pathogens during the outbreak, and who wants to do that? Another issue is to choose where and when these seeds should appear. Once again, no rationale is both simple and consistent with basic facts about real epidemics. Yet an issue with using many seeds is that they would make the outbreak happen with a much higher chance than one seed, which thus takes the edge off a very characteristic feature of all compartmental models—the early die-outs. If one is modeling bioterrorism, having many seeds could make sense, otherwise avoid it.
- Pick the source node uniformly at random. This is in the spirit of doing things as random as possible, given the available information. Probably there is a correlation between, say, the degree and the probability of being a source of an outbreak, but we don’t know it, so let’s do it like this.
- Infect the source at a time chosen randomly (with uniform randomness between the beginning and end of the temporal network). This is also to do things as random as possible. Clearly, it would be wrong to choose a time relating to features of the data (like the beginning of the data, or at the time when an individual enters the data). We did that anyway in some papers, and probably it doesn’t affect the conclusions much, but these are very biased time points. For example, in many data sets of human activity, individuals behave differently at the beginning of their presence in the data, and the end. This choice means that outbreaks starting towards the end will not have the time to spread to the entire network, which is bad news if one wants to study the propagations of large outbreaks (I usually don’t). Otherwise, I can’t see any methodological problem with it.
- Not make a node infectious at the time step when it gets infected. In empirical data sets, nodes typically have many simultaneous contacts. This rule is to prohibit the infection from spreading further than one step at a time. I.e. if there are contacts (A,B,10) and (B,C,10), but no (A,C,10), then A cannot infect C (via B) at time 10. So technically, our model is an SEIR model with the E stage lasting an infinitesimal time. Optionally (and what seemed more principled to me before), one can assume that the contacts and infection processes are truly instantaneous but happen in a specific order. Then one needs to average over randomizations over the order of the contacts during a time step.
- Let the infection probability be the same for all contacts. This is a well-known oversimplification, but I do it anyway. Partly to conform to conventions, partly because exploring realistic distributions of infection probabilities and infectiousness is a bit boring topic (but sure, eventually I’ll get there . . ).
- Let the infection last an exponentially distributed time. This is to conform to the literature and be able to compare results to analytical results for static graphs. Many times, I have used a fixed time of the infection duration. This is not realistic either, and it gives some spurious resonance-like effects with cyclic temporal patterns in empirical data.
