June 21, that authors of the General Network Dismantling paper sent me a reply with their comments on this blog post. You can read it here. I will comment on it later.
This post is about some recent experiences and thoughts of reproducing the computational results of a paper.
Thoughts about computational reproducibility
The reproducibility of results in computational science has become a hotter topic over the last few years. I have to admit that I haven’t followed the discussion very closely. I know that some journals and societies have guidelines, but for this blog post, I will not discuss these, just share my opinions and experiences.
What are the code-writing authors’ duties? To write correct and readable code that is fully described in the paper, and to share at least the core of it. (I think code to plot figures, generate batch scripts, do simple post-processing, and code that is very standard or short shouldn’t be shared—that’s just clutter.) They should also take time to answer questions about their code, etc.
…the co-authors’ duties? It’s tempting to say all authors should share the responsibility equally, but that wouldn’t reflect the way research groups actually work. On the other hand, (if there are co-authors), it makes no sense if no one other than the coder tries to implement the code and test it. Pair programming is not a bad idea, either. This is probably the trickiest of the responsibilities to judge. In some scenarios, a lab could have a too-pushy atmosphere that makes coders under check their programs, then isn’t it the lab leader’s fault? Etc.
… reviewers’ duties? To try to implement the code, and check it closer if it seems suspicious.
… journals’ duties? To not accept manuscripts with code red-flagged by reviewers. Swiftly handle raised concerns about published papers. Even if conclusions are unaffected, they should publish errata with correct figures. Promote the sharing of code.
… readers’ duties? Not much, unless they need to implement the code. If they do and discover problems, they should share their concerns. I think this must be recognized as an essential community service and merit because it takes time and risks to create tensions with the authors. I do feel reluctant to report my reproduction attempts (below), but if I don’t, then others need to spend time, and I don’t think it would benefit anyone in the end.
Assuming a paper is based on erroneous code, what should be the consequences? That is not an easy yes or no question—options from doing nothing to retraction are all possible. Of course, the bolder the claims, and the more prestigious the journal, the higher the demands on correct code should be. Their grey zone is also vast. If the description of the algorithm is ambiguous (which is a widespread case), then who has the right to interpret it? It should preferably be an open dialogue.
Generalized network dismantling
On to my own recent experience. I have some research interests in network dismantling (a.k.a. network attack, network fragmentation, etc.)—how to delete n nodes from a network to decrease the size of the largest connected component as much as possible. This is an NP-hard problem, so the name of the game is to come up with heuristics. I’m not designing heuristics myself, but I need to have a collection of them, so that’s my motivation to follow this development.
A recent paper in PNAS “Generalized network dismantling” by Ren, Gleinig, Helbing, and Antulov-Fantulina does just that—proposes a new algorithm. They furthermore relax the constraint that deleting one node costs one unit—in general, it could be any function that maps a node in a network to a number. They test their algorithm for the cases where this function is the degree of the node (and the standard, unit-cost case). The algorithm generalizes the classical method by Fiedler to partition graphs by the sign of the eigenvector corresponding to the second smallest eigenvalue of the Laplacian. (A bit interesting, because I thought that was considered just a historical curiosity.) They: 1. Construct a similar spectral partitioning for the general case. 2. Find the cheapest set of nodes to delete to carry out the partitioning. 3. Repeat.
Random numbers
The first reason I was puzzled by the authors’ C++ implementation (available on github), was that I didn’t understand that the unit-cost case needed some hacking as the paper almost only talks about the degree-cost case. That was more or less my own stupidity, but as I looked at the code, I noticed that they initialized the random number generator to the default seed (1) at every call of the function, partitioning the graph. In other words, they did not really use random numbers, at least not as the praxis is in the field. After fixing that, the output started depending on the seed. Here is the figure (Fig. S5) that I aimed to reproduce. The (GND) line of the paper is (in black), and 10 runs of my code with the handling of random numbers fixed (in color).
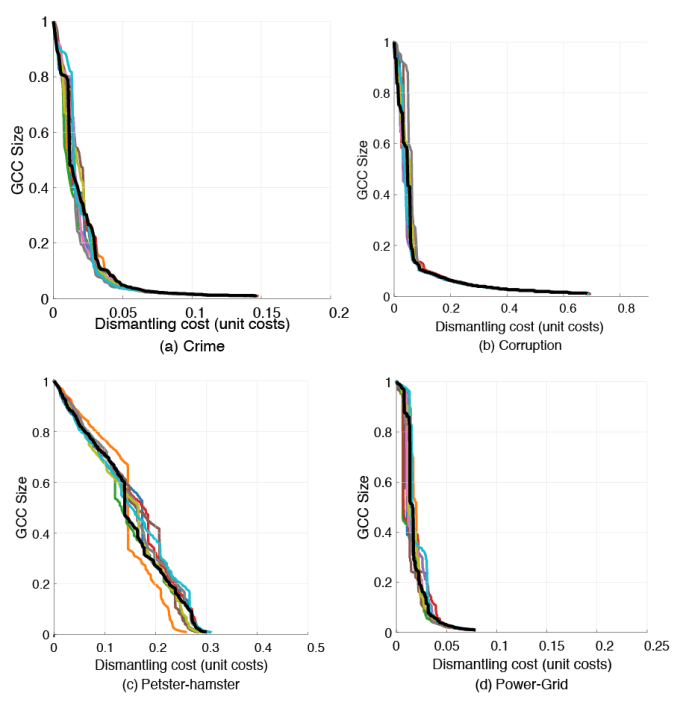
But the algorithm itself is almost deterministic. It makes sense to use randomness to break ties in the deletion process so that the output is independent of the labeling of the nodes, but the authors’ code doesn’t do that, and it would not produce such a large variation. So what more is wrong?
Eigenvectors
It turns out that the authors solve the eigenvector problem not by calling a linear-algebra library routine, but by their own power iteration. The core of linear algebra libraries (BLACS) is optimized to an incredible level. People in the CS department of my alma mater were into that, and I almost don’t exaggerate if I say theses were written about how to shave off a couple of scalar multiplications per matrix multiplication. (Also, the way they minimize the cache loads is pretty fantastic.) Then for solving eigenvalue problems, there is another layer of close-to-magical code (all in Fortran, of course), so why would you want to write your own eigenvector solver?
The problem with the authors’ code is that they use power iterations (an often slow method) with a fixed number of iterations rather than a standard convergence criterion. Here is a figure of how the Hamming distance of the sign of the eigenvectors (the y-axis) converges in their code (this is the first partition of the “Crime_Gcc” data set and the degree-cost case). The red cross marks where the authors’ code breaks the iterations.

Simply running the iterations 500 times longer makes the curves stable again, but different from the paper:
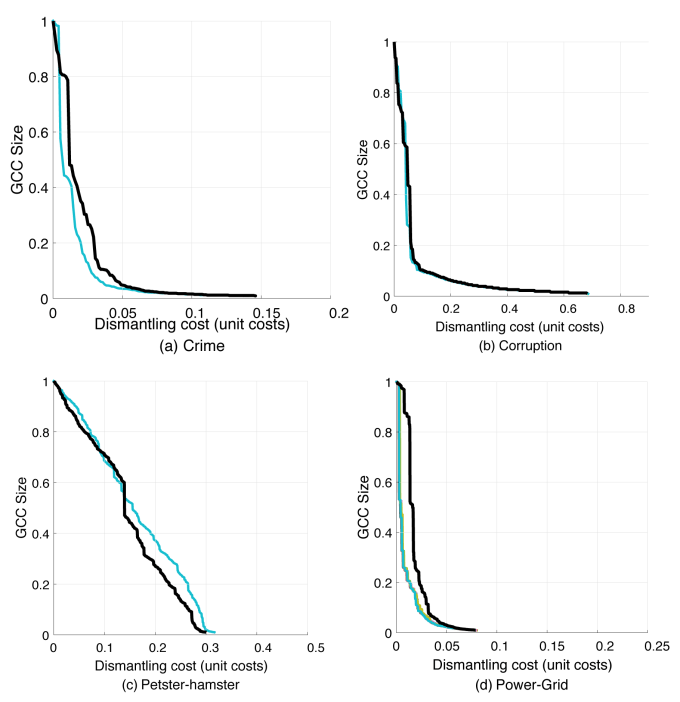
My own code
At this point, I thought it was best to code it from scratch. Since the eigenvector problem is the bottleneck, there is not much to gain in coding it in C++, I thought, so I wrote it in Python. My goal was to implement it as close to the description in the paper as I could. Ultimately, I just want a working implementation of the method and be able to point to the article for a description. You can get the code here. Feel free to check it and tell me if I got something wrong.
This is the output of my code vs. the curves from the paper.
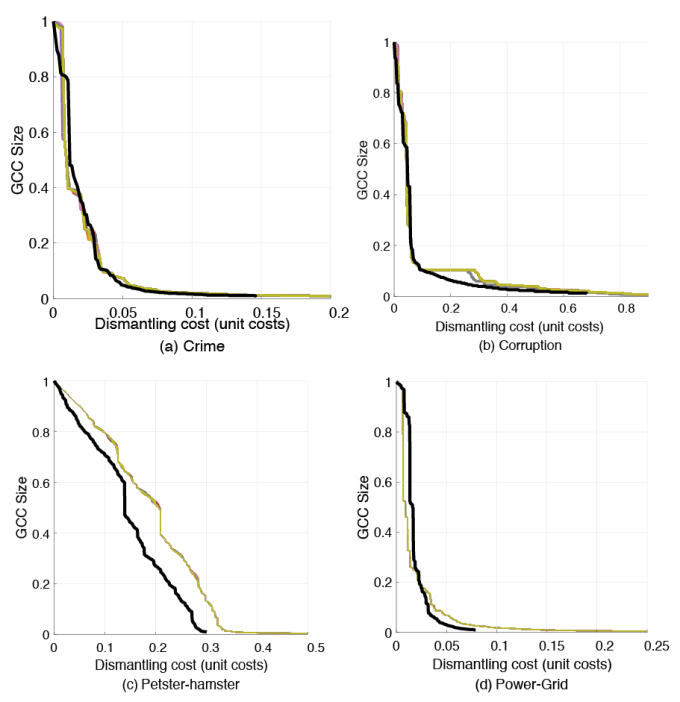
Note that there is still some randomness from the fact that I use randomness to break ties (as mentioned above).
The authors’ code output nodes of the vertex cover set (step 2 above) in order of decreasing degree (for the degree-weight case) or increasing degree (for the unit-weight case). This is a bit curious because one can no longer get the unit-degree case by replacing degrees with one somewhere in the algorithm (so it is no longer generalized network dismantling). If I also do that, the spread of the curves above goes away. I don’t know where the deviations from my code and the converged curves of the authors’ code come from, but I feel my time for checking their code is up.
Concluding thoughts
Although the curves of the paper do not seem to represent the described algorithm, I think the conclusions of the articles are correct. I also like the ideas of the algorithm a lot and think the paper is a valuable contribution. Of course, I’m a bit disappointed that this took a lot of my time, but I still respect the authors very much and look forward to their future work. I use other results from the same authors with nothing to complain about. It would, however, be good to see all figures of the paper generated with the correct code.